|
4/8/2023 0 Comments Ribosomal insidia bacteria The eukaryotic 60S subunit structure was also determined from T. The complete structure of a eukaryotic 40S ribosomal structure in Tetrahymena thermophila was published and described, as well as much about the 40S subunit's interaction with eIF1 during translation initiation. Ribosome structures at atomic resolution in the 1990s, it took another decade until in 2011, high resolution structures of eukaryotic ribosome were obtained by X-ray crystallography, mainly because of the difficulties in obtaining crystals of sufficient quality. Īfter the determination of the first bacterial More recently structures at sub-nanometer resolution were obtained for complexes of ribosomes and factors involved in translation. Higher resolution structures of the yeast ribosome by cryo-electron microscopy allowed the identification of protein and RNA structural elements. Initial structures of eukaryotic ribosomes were determined by electron microscopy.įirst 3D structures were obtained at 30–40 Å resolution for yeast Recent research suggests heterogeneity in the ribosomal composition, i.e., that the stoichiometry among core ribosomal proteins in wild-type yeast cells and embryonic stem cells depends both on the growth conditions and on the number of ribosomes bound per mRNA. For a detailed list of proteins, including archaeal and bacterial homologs please refer to the separate articles on the 40S and 60S subunits. In addition, it contains a 5.8S rRNA that corresponds to the 5' end of the 23S rRNA, and a short 5S rRNA.īoth 18S and 28S have multiple insertions to the core rRNA fold of their prokaryotic counterparts, which are called expansion segments. The 60S subunit contains a 28S rRNA that is homologous to the prokaryotic 23S ribosomal RNA. The 40S subunit contains a 18S ribosomal RNA (abbreviated 18S rRNA), which is homologous to the prokaryotic 16S rRNA. Furthermore, several additional proteins are found in the small and large subunits of eukaryotic ribosomes, which do not have prokaryotic homologs. The small subunit monitors the complementarity between tRNA anticodon and mRNA, while the large subunit catalyzes peptide bond formation.Ĭompared to their prokaryotic homologs, many of the eukaryotic ribosomal proteins are enlarged by insertions or extensions to the conserved core. Both subunits contain dozens of ribosomal proteins arranged on a scaffold composed of ribosomal RNA (rRNA). Eukaryotic ribosomes have two unequal subunits, designated small subunit (40S) and large subunit (60S) according to their sedimentation coefficients. Eukaryotic ribosomes are also known as 80S ribosomes, referring to their sedimentation coefficients in Svedberg units, because they sediment faster than the prokaryotic ( 70S) ribosomes. However, the ribosomes of eukaryotes (animals, plants, fungi, and large number unicellular organisms all with a nucleus) are much larger than prokaryotic ( bacterial and archaeal) ribosomes and subject to more complex regulation and biogenesis pathways. Ribosomes from all organisms share a highly conserved catalytic center. The ribosome selects aminoacylated transfer RNAs (tRNAs) based on the sequence of a protein-encoding messenger RNA (mRNA) and covalently links the amino acids into a polypeptide chain. Ribosomes are a large and complex molecular machine that catalyzes the synthesis of proteins, referred to as translation. PDB identifiers 4a17, 4A19, 2XZM aligned to 3U5B, 3U5C, 3U5D, 3U5E Proteins shared only between eukaryotes and archaea are shown in orange, and proteins specific to eukaryotes are shown in red. These proteins have homologs in eukaryotes, archaea and bacteria. 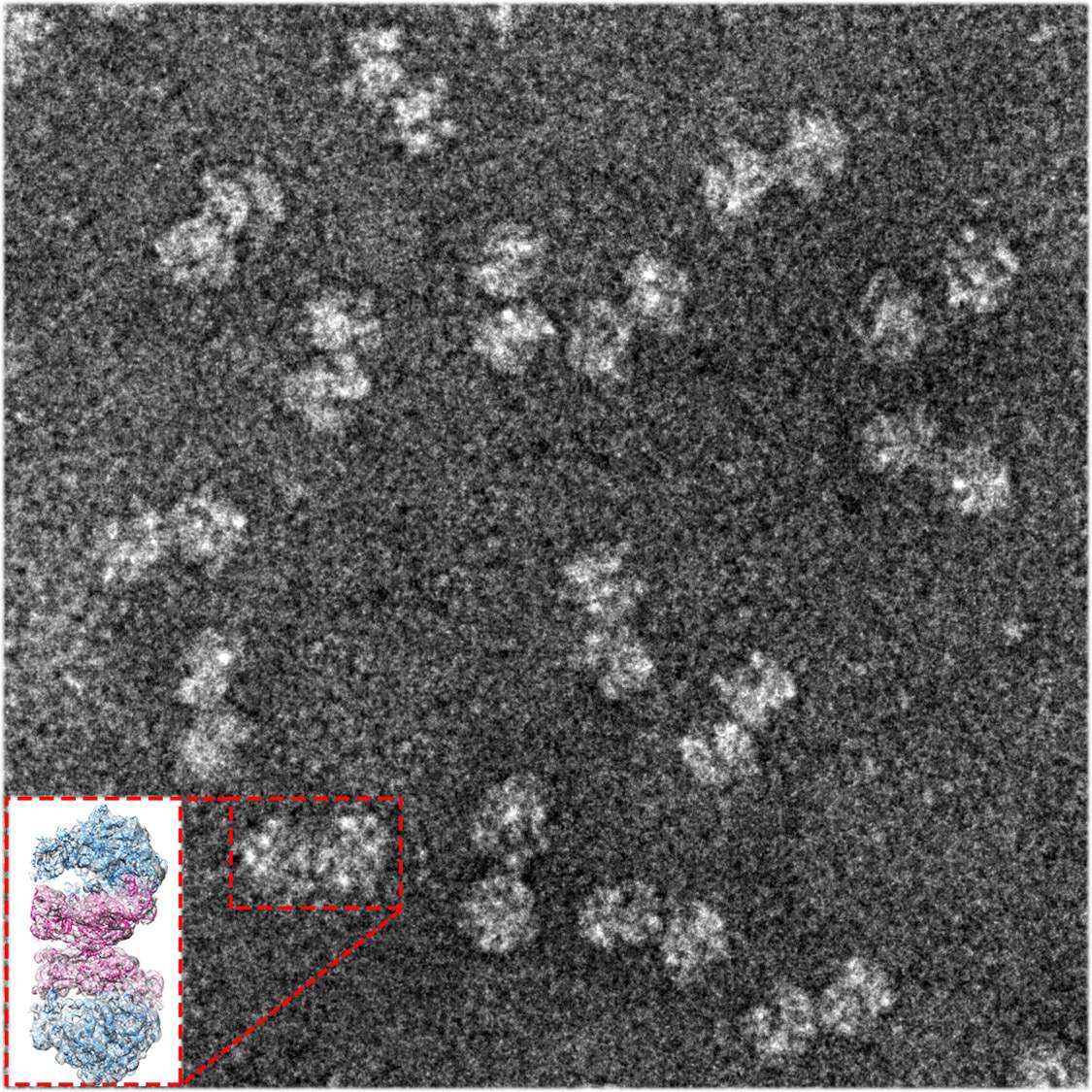
Universally conserved proteins are shown in blue. 
The ribosomal RNA ( rRNA) core is represented as a grey tube, expansion segments are shown in red. The 40S subunit is on the left, the 60S subunit on the right. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
